Honorable Mention
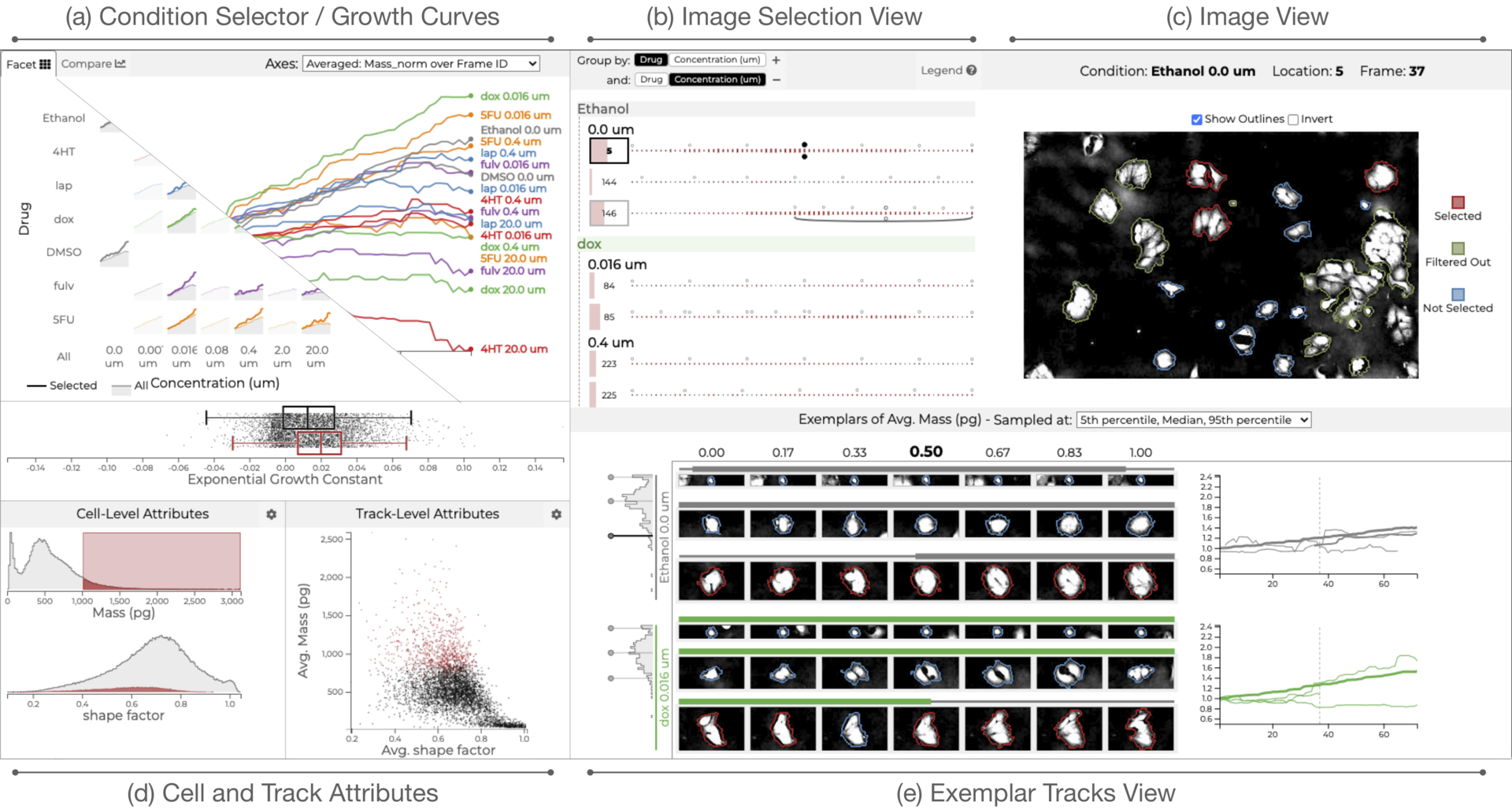
Loon: Using Exemplars to Visualize Large Scale Microscopy Data
Devin Lange, Eddie Polanco, Robert Judson-Torres, Thomas Zangle, Alexander Lex
External link (DOI)
View presentation:2021-10-29T15:30:00ZGMT-0600Change your timezone on the schedule page
2021-10-29T15:30:00Z
Fast forward
Direct link to video on YouTube: https://youtu.be/iRsL3WiZbhI
Abstract
Which drug is most promising for a cancer patient? This is a question a new microscopy-based approach for measuring the mass of individual cancer cells treated with different drugs promises to answer in only a few hours. However, the analysis pipeline for extracting data from these images is still far from complete automation: human intervention is necessary for quality control for preprocessing steps such as segmentation, to adjust filters, and remove noise, and for the analysis of the result. To address this workflow, we developed Loon, a visualization tool for analyzing drug screening data based on quantitative phase microscopy imaging. Loon visualizes both derived data such as growth rates, and imaging data. Since the images are collected automatically at a large scale, manual inspection of images and segmentations is infeasible. However, reviewing representative samples of cells is essential, both for quality control and for data analysis. We introduce a new approach of choosing and visualizing representative exemplar cells that retain a close connection to the low-level data. By tightly integrating the derived data visualization capabilities with the novel exemplar visualization and providing selection and filtering capabilities, Loon is well suited for making decisions about which drugs are suitable for a specific patient.